Baird RB, Hitchcock TJ, Ševčik J, Monteith KM, Gardner A, Ross L & Mongue AJ (2025) Faster adaptation but slower divergence of X chromosomes under paternal genome elimination. Nature Communications 16, 5288.
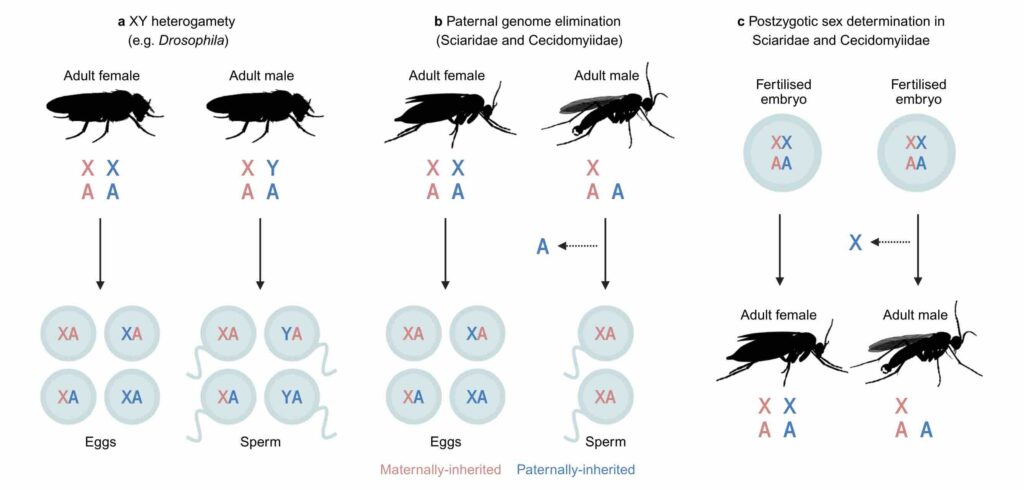
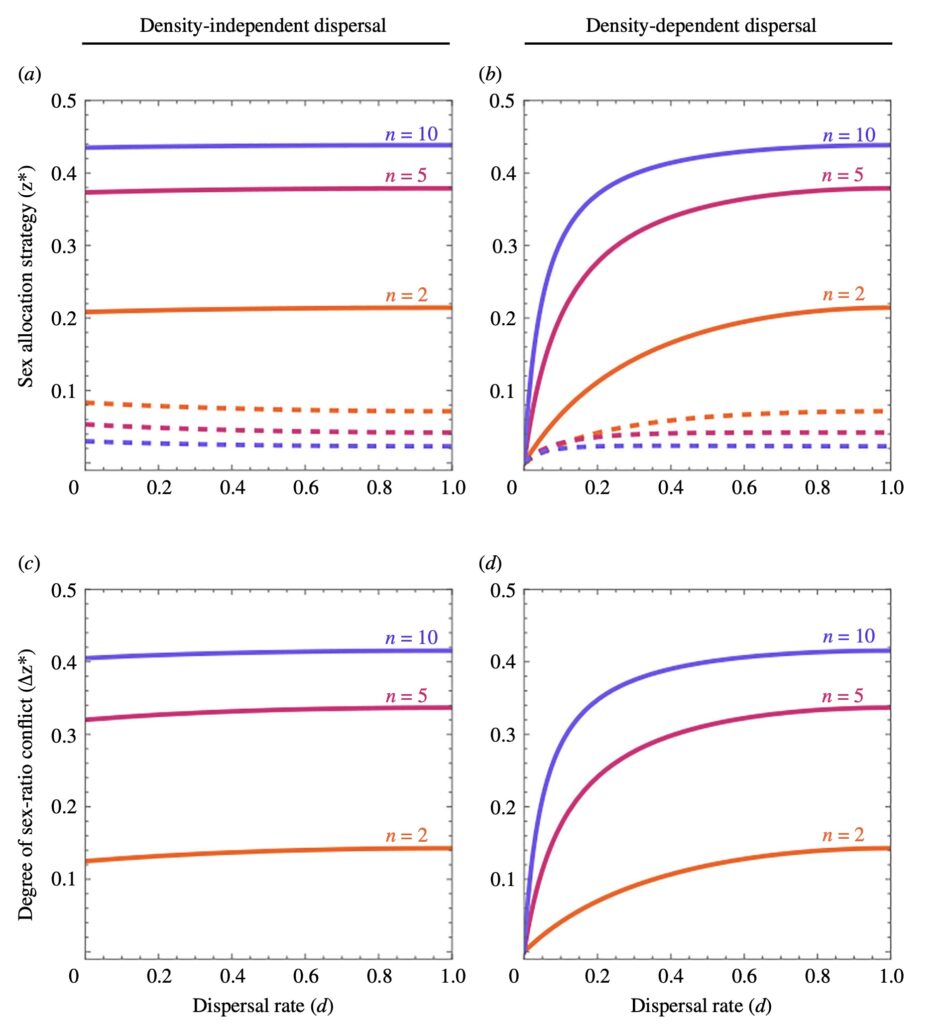
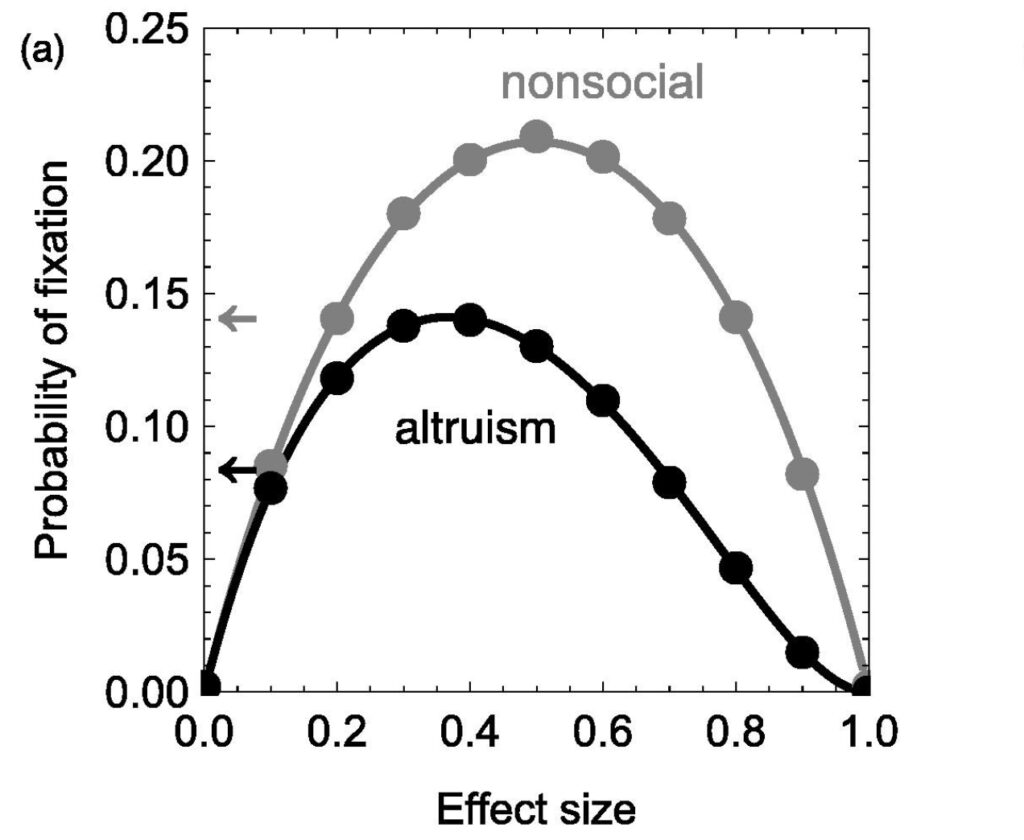
Differences in transmission and ploidy between sex chromosomes and autosomes drive divergent evolutionary trajectories, with sex chromosomes generally evolving faster. Because sex-linked genes are transmitted less frequently, they are under less efficient selection. Conversely, exposure of recessive mutations on haploid sex chromosomes creates more efficient selection. In most systems, these effects occur simultaneously and are confounded. The fly families Sciaridae (fungus gnats) and Cecidomyiidae (gall midges) have X0 sex determination, but males transmit only maternally inherited chromosomes. This phenomenon results in equal transmission of the X and autosomes, allowing the effect of haploid selection to be studied in isolation. We discover that, unlike well-studied systems, X chromosomes diverge more slowly than autosomes in these flies. Using population genomic and expression data, we show that despite the X evolving more adaptively, stronger purifying selection explains slower divergence. Our findings demonstrate the utility of non-Mendelian inheritance systems for understanding fundamental evolutionary processes.